Chapter 3 of 8
Searching for articles
A core early step is deciding which literature databases to search. That choice should follow the specialty and scope of the review: clinical, methods-focused, multidisciplinary, and emerging fields each lean on different combinations of sources.
In AIPRA, you attach each literature source you plan to search as a database on the project. The interface lists the sources currently available to add and configure for your review.
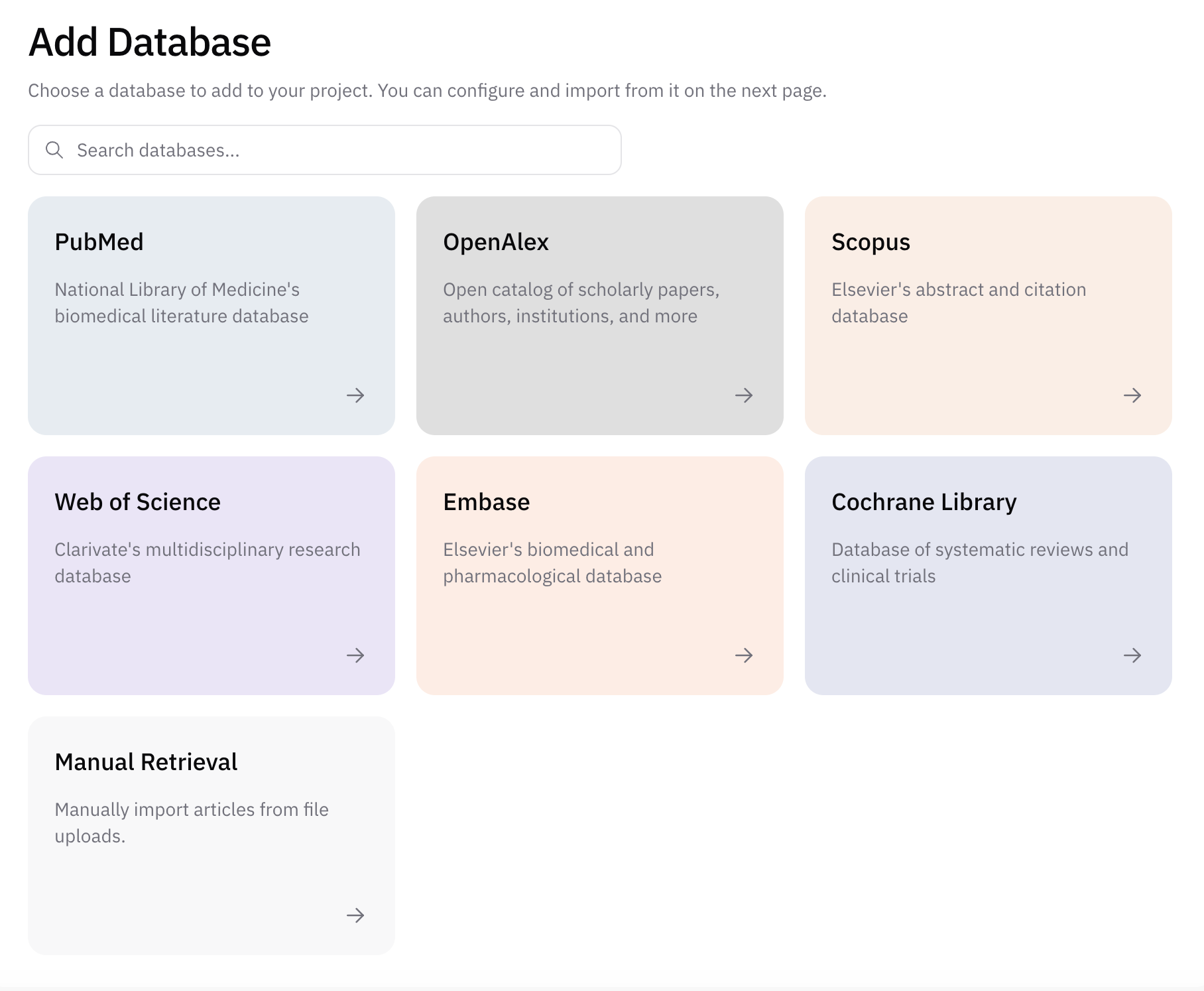
The interactive reference below summarizes six databases teams often consider in the wider literature. Use the pills to jump to a source, or expand a row for full detail on volume, coverage, access, and technical notes.
Common literature databases
Choose a database to see scope, access, and technical notes. Your specialty and question should drive which sources you prioritize; many reviews search at least two independent bibliographic databases.
| Database | Scale | Access | |
|---|---|---|---|
| 37M+ citations | Free / open | ||
| |||
| ~474M works | Free / open | ||
| |||
| 90M+ records | Subscription | ||
| |||
| 170M+ (network) | Tiered | ||
| |||
| 40M+ records | Subscription | ||
| |||
| 9k+ reviews; 2M+ CENTRAL | Free / open | ||
| |||
You can also document and add results from any additional database not listed here when your protocol requires it.
Practical selection guidance
A common recommendation is to include at least two independent bibliographic databases so retrieval bias is reduced. For example, in many medical reviews, teams pair PubMed with Embase or Web of Science (depending on institutional access and the clinical question).
OpenAlex is especially useful when you want broad, multidisciplinary coverage and easier programmatic access. It also surfaces a large share of gray literature—for example preprints that are not yet formally published in a peer-reviewed journal—which can matter for fast-moving topics (such as work on large language models) where delaying until full publication would miss important evidence.
From concepts to a search query
Each database expects searches to be expressed in its own syntax and field rules. That expression is your search query. Building one is usually a multistep process.
- Identify the main concepts implied by the research question. For example, for “The use of AI in diabetic retinopathy screening,” core concepts might be: AI, diabetic retinopathy, and screening.
- Expand each concept with synonyms and related terms that authors might have used in titles, abstracts, or keywords. The goal is sensitivity: retrieve relevant studies even when terminology differs. For the AI concept, examples include: machine learning (ML), automated systems, automation, neural networks, deep learning, natural language processing (NLP), cognitive computing, intelligent agents, computer vision, and knowledge representation—always tailored to what fits your question and database.
- Assemble the database-specific query using those terms, Boolean operators, proximity or phrase rules, and field limits as required by each platform (for example MeSH in PubMed versus Emtree in Embase).
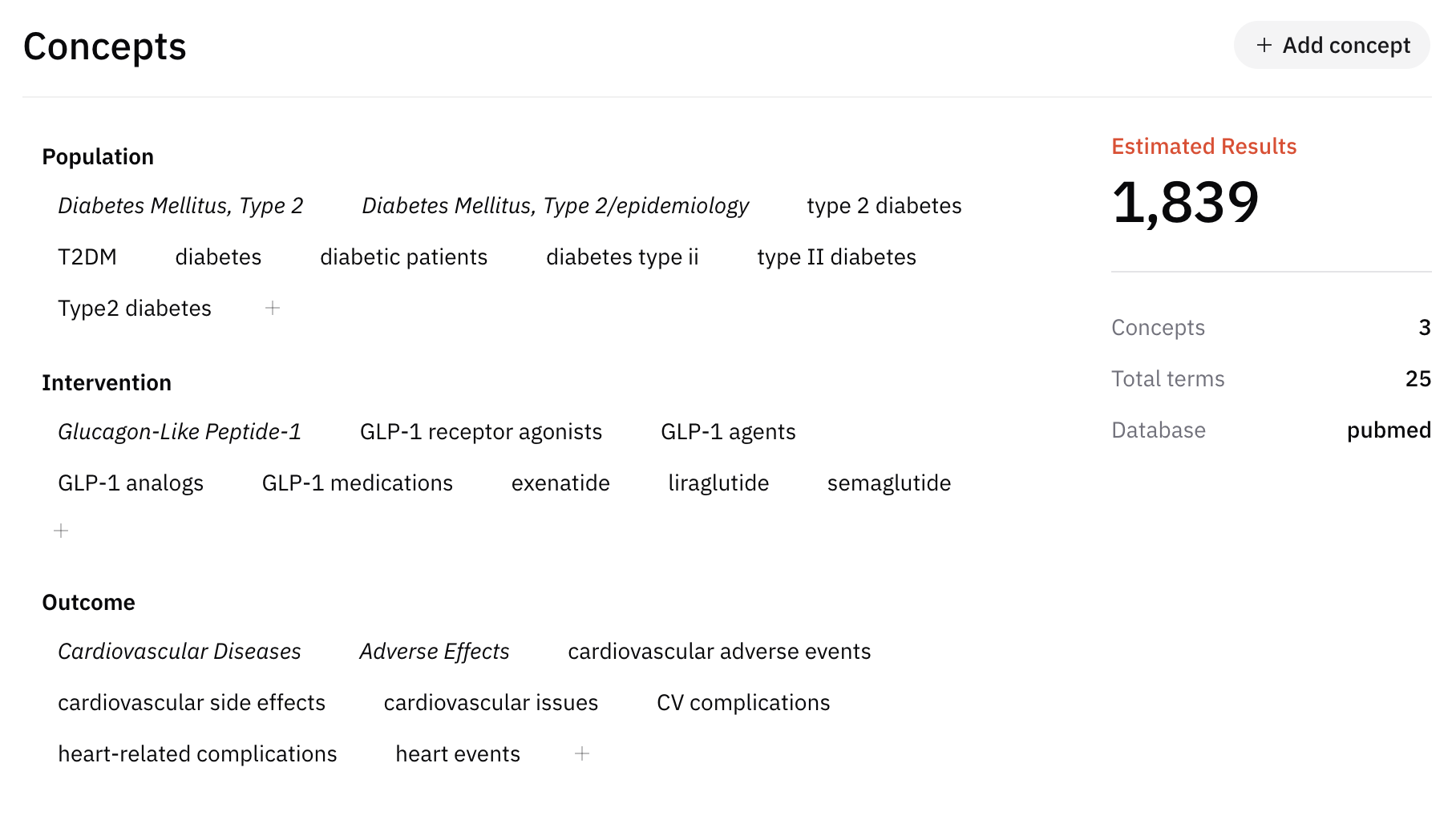
How AIPRA helps
AIPRA can suggest concepts and synonym lists starting from your research question. When you select a database in the workflow, it can also help draft a query aligned with that source’s typical syntax— which you should still validate against the live database and your information specialist’s advice.
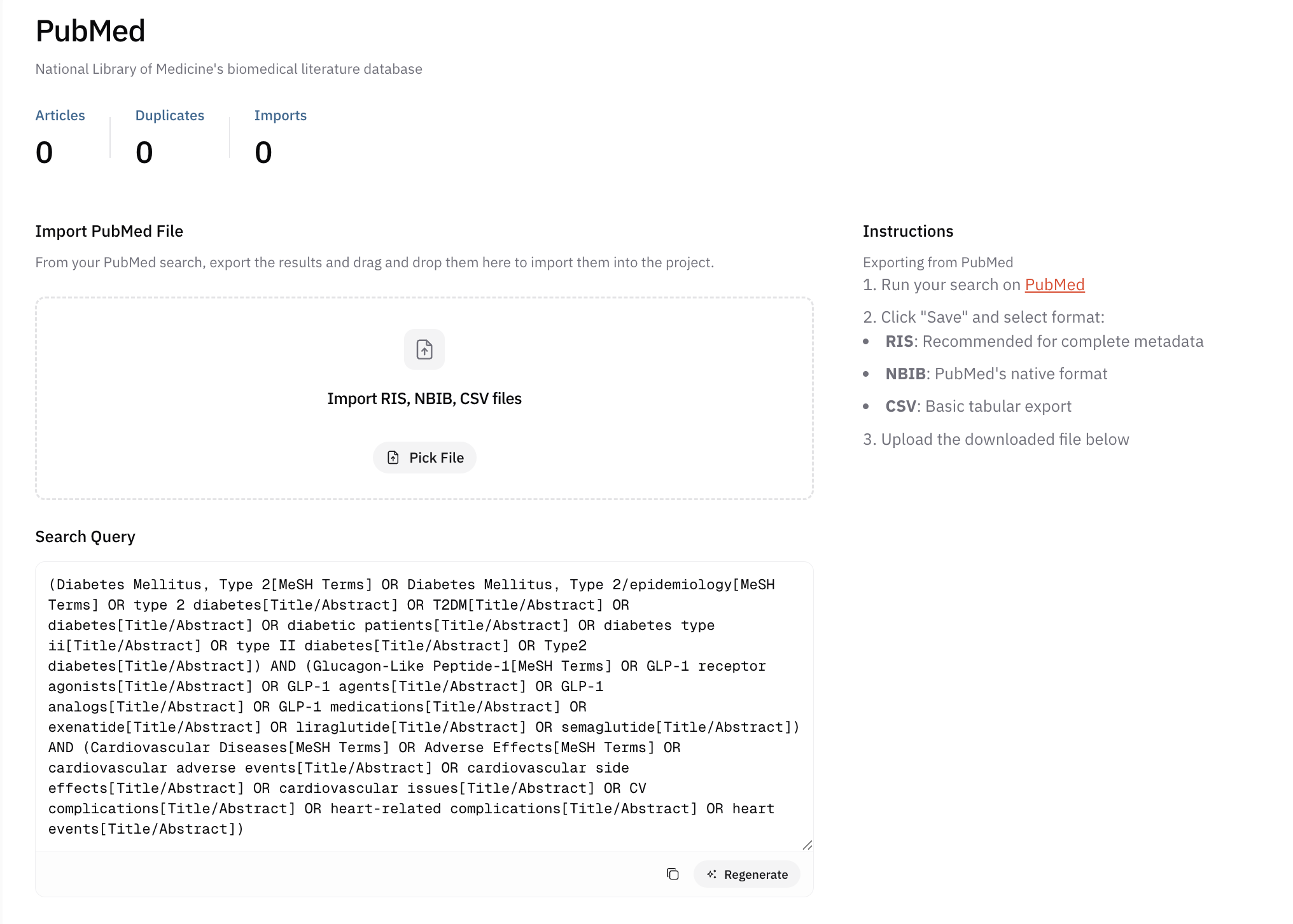
In practice, you will usually run the search yourself in each database’s native interface: open the site, paste or adapt the query copied from AIPRA, run the search, export the records your protocol specifies, and upload those exports into AIPRA’s database area for the project so screening and deduplication stay centralized.
Deduplication across databases
Searching multiple databases almost always retrieves the same article more than once when it is indexed in several places. AIPRA runs a deduplication pass using unique identifiers such as DOI and PMID, which catches most duplicate records automatically.
Identifier-based matching is strong but not perfect; a small number of duplicates may still appear during screening (for example when metadata differs between exports). Teams should plan to resolve those remaining duplicates as part of title/abstract or full-text review.
For methodological context on deduplication in systematic reviews, see the Cochrane Handbook section on deduplication: Cochrane Handbook — duplicate reports and deduplication.